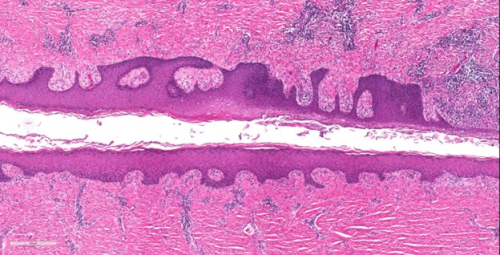
Genetic Variants Associated with Hidradenitis Suppurativa
August 27, 2023
Paper Citation: Sun Q, Broadaway KA, Edmiston SN, et al. Genetic Variants Associated With Hidradenitis Suppurativa. JAMA Dermatol. Published online July 26, 2023. doi:10.1001/jamadermatol.2023.2217
Plain Language Summary by: Linnea Westerkam BA1, Christopher Sayed MD2
1University of North Carolina School of Medicine
2University of North Carolina Department of Dermatology
Why was this study done?
About half of patients with hidradenitis suppurativa (HS) have a family member that’s also affected. For many people there may be something in their genes that puts them at risk of ultimately developing HS. This study was done to try to identify changes in genes that might be associated with HS.
How was the study done?
720 patients with HS were recruited at the University of North Carolina. Genetic studies, called genome-wide association studies or GWAS, were performed and analyzed alongside three other databases with many more patients with HS. Their DNA was compared to the DNA of many thousands of people without HS to find differences unique to people with HS.
What did the study find?
Changes in the DNA of HS patients near 2 genes calledSOX9 and KLF5 were identified that are likely associated with HS. In patients with HS, inflammation likely centers around hair follicles. SOX9 is important for hair follicle development and in mouse studies, mice without SOX9 developed breakdown of hair follicles and scarring similar to some of the features of HS. KLF5 is important for skin development associated with other skin conditions (atopic dermatitis, commonly known as eczema, and non-healing wounds) and colitis (gut inflammation). Since these genes affect skin and hair development and local inflammation, they likely impact the risk of developing HS. They may also play a role in disease progression.
What are next steps?
Additional studies are needed to confirm the impact of these genetic changes on how HS disease develops and how severe it becomes. We also need to understand how the changes discovered in the DNA affect how these genes work in HS. To identify other rarer genetic variants associated with HS, larger studies are necessary. These results will hopefully lead to further research, improve understanding of HS, and help us learn how to develop new treatments for HS.
Image credit: Paul Googe, MD

